Posts tagged DNA
CU scientists find life forms in a lifeless land
Jun 14th
A new DNA analysis of rocky soils in the Martian-like landscape on some volcanoes in South America has revealed a handful of bacteria, fungi and other rudimentary organisms called archaea, which seem to have a different way of converting energy than their cousins elsewhere in the world.

“We haven’t formally identified or characterized the species,” said Ryan Lynch, a CU-Boulder doctoral student involved in the study. “But these are very different than anything else that has been cultured. Genetically, they’re at least 5 percent different than anything else in the DNA database of 2.5 million sequences.”
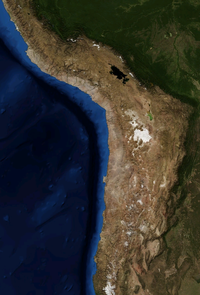
Life gets little encouragement on the incredibly dry slopes of the tallest volcanoes in the Atacama region, where CU-Boulder Professor Steve Schmidt and his team collected soil samples. Much of the sparse snow that falls on the terrain sublimates back to the atmosphere soon after it hits the ground, and the soil is so depleted of nutrients that nitrogen levels in the scientists’ samples were below detection limits.
One of the most hostile environments on the planet
Ultraviolet radiation in the high-altitude environment can be twice as intense as in a low-elevation desert, said Schmidt of CU-Boulder’s ecology and evolutionary biology department. While the researchers were on site, temperatures dropped to 14 degrees Fahrenheit one night and spiked to 133 F the next day.
How the newfound organisms survive under such circumstances remains a mystery. Although Ryan, Schmidt and their colleagues looked for genes known to be involved in photosynthesis and peered into the cells using fluorescent techniques to look for chlorophyll, they couldn’t find evidence that the microbes were photosynthetic.
Instead, they think the microbes might slowly generate energy by means of chemical reactions that extract energy and carbon from wisps of gases such as carbon monoxide and dimethylsulfide that blow into the desolate mountain area. The process wouldn’t give the bugs a high-energy yield, Lynch said, but it could be enough as it adds up over time. A paper on the findings has been accepted by the Journal of Geophysical Research-Biogeosciences, published by the American Geophysical Union.
While normal soil has thousands of microbial species in just a gram of soil, and garden soils even more, remarkably few species have made their home in the barren Atacama mountain soil, the new research suggests. “To find a community dominated by less than 20 species is pretty amazing for a soil microbiologist,” Schmidt said.
Nearly 20,000 feet in altitude, snowless for 48,000 years
He has studied sites in the Peruvian Andes where, four years after a glacier retreats, there are thriving, diverse microbe communities. But on these volcanoes on the Chile-Argentina border, which rise to altitudes of more than 19,685 feet and which have been ice-free for 48,000 years, the bacterial and fungal ecosystems have not undergone succession to more diverse communities. “It’s mostly due to the lack of water, we think,” he said. “Without water, you’re not going to develop a complex community.”
“Overall, there was a good bit lower diversity in the Atacama samples than you would find in most soils, including other mountainous mineral soils,” Lynch said. That makes the Atacama microbes very unusual, he added. They probably had to adapt to the extremely harsh environment, or may have evolved in different directions than similar organisms elsewhere due to long-term geographic isolation.
Growth on the mountain might be intermittent, Schmidt suggested, especially if soils only have water for a short time after snowfall. In those situations, there could be microbes that grow when it snows, then fall dormant, perhaps for years, before they grow again. High-elevation sites are great places to study simple microbial communities, ecosystems that haven’t evolved past the very basics of a few bacteria and fungi, Schmidt said.
“There are a lot of areas in the world that haven’t been studied from a microbial perspective, and this is one of the main ones,” he said. “We’re interested in discovering new forms of life, and describing what those organisms are doing, how they make a living.”
Schmidt’s lab, along with others, is studying how microorganisms travel from one site to another. One common method of microbe transport is through the air — they’re caught up in winds, sucked up into clouds, form rain droplets and then fall back to the ground somewhere else as precipitation.

But on mountains like Volcán Llullaillaco and Volcán Socompa, the high UV radiation and extreme temperatures make the landscape inhospitable to outside microbes. “This environment is so restrictive, most of those things that are raining down are killed immediately,” Schmidt said. “There’s a huge environmental filter here that’s keeping most of these things from growing.”
The next steps for the researchers are laboratory experiments using an incubator that can mimic the extreme temperature fluctuations to better understand how any organism can live in such an unfriendly environment. Studying the microbes and finding out how they can live at such an extreme can help set boundaries for life on Earth, Schmidt said, and tells scientists what life can stand. There’s a possibility that some of the extremophiles might utilize completely new forms of metabolism, converting energy in a novel way.
Schmidt also is working with astrobiologists to model what past conditions were like on Mars. With their rocky terrain, thin atmosphere and high radiation, the Atacama volcanoes are some of the most similar places on Earth to the Red Planet.
“If we know, on Earth, what the outer limits for life were, and they know what the paleoclimates on Mars were like, we may have a better idea of what could have lived there,” he said.
Other paper authors included Andrew King of Ecosystem Sciences, CSIRO Black Mountain in Acton, Australia; Mariá Farías of Laboratorio de Investigaciones Microbiologicas de Lagunas Andinas, Planto Piloto de Procesos Industriales Microbiologicas, CCT, CONICET in Tucuman, Argentina; Preston Sowell of Geomega, an environmental consulting firm in Boulder; and Christian Vitry of Museo de Arqueologia de Alta Montana in Salta, Argentina.

NEW CU-BOULDER STUDY REVEALS BACTERIA FROM DOG FECES IN OUTDOOR AIR OF URBANIZED AREAS
Aug 18th
Bacteria from fecal material — in particular, dog fecal material — may constitute the dominant source of airborne bacteria in Cleveland’s and Detroit’s wintertime air, says a new University of Colorado Boulder study.
The CU-Boulder study showed that of the four Midwestern cities in the experiment, two cities had significant quantities of fecal bacteria in the atmosphere — with dog feces being the most likely source.
“We found unexpectedly high bacterial diversity in all of our samples, but to our surprise the airborne bacterial communities of Detroit and Cleveland most closely resembled those communities found in dog poop,” said lead author Robert Bowers, a graduate student in CU-Boulder’s ecology and evolutionary biology department and the CU-headquartered Cooperative Institute for Research in Environmental Sciences, or CIRES. “This suggests that dog poop may be a potential source of bacteria to the atmosphere at these locations.”
The study was published July 29 in Applied and Environmental Microbiology. Co-authors on the study included Noah Fierer, an assistant professor in CU-Boulder’s ecology and evolutionary biology department and a CIRES fellow; Rob Knight, an associate professor in CU-Boulder’s chemistry and biochemistry department; Amy Sullivan and Jeff Collett Jr. of Colorado State University; and Elizabeth Costello of the Stanford University School of Medicine.
Scientists already knew that bacteria exist in the atmosphere and that these bacteria can have detrimental effects on human health, triggering allergic asthma and seasonal allergies, Fierer said. But it is only in recent years that researchers have realized that there is an incredible diversity of bacteriaresiding in the air, he said.
“There is a real knowledge gap,” said Fierer. “We are just starting to realize this uncharted microbial diversity in the air — a place where you wouldn’t exactly expect microbes to be living.”
To gain further understanding of just what microbes are circulating in urban environments, the team analyzed the local atmosphere in the summer and winter at four locations in the Great Lakes region of the U.S. Three of the locations — Chicago, Cleveland and Detroit — are major cities with populations of greater than 2 million, and one location, Mayville, Wis., is a small town with a population of less than 6,000.
The team used nearly 100 air samples collected as part of a previous study conducted by Colorado State University. The CSU experiment investigated the impact of biomass burning and involved studying the impacts of residential wood burning and prescribed fires on airborne fine particle concentrations in the midwestern United States.
“What we’ve been looking at are the numbers and the types of bacteria in the atmosphere,” Fierer said. “We breathe in bacteria every minute we are outside, and some of these bugs may have potential health implications.”
The researchers analyzed the bacteria’s DNA in the collected air samples and compared the bacteria they found against a database of bacteria from known sources such as leaf surfaces, soil, and human, cow and dog feces. They discovered that the bacterial communities in the air were surprisingly diverse and also that, in two of the four locations, dog feces were a greater than expected source of bacteria in the atmosphere in the winter.
In the summer, airborne bacteria come from many sources including soil, dust, leafsurfaces, lakes and oceans, Bowers said. But in the winter, as leaves drop and snow covers the ground, the influence that these environments have as sources also goes down. It is during this season that the airborne communities appeared to be more influenced by dog feces than the other sources tested in the experiment, he said.
“As best as we can tell, dog feces are the only explanation for these results,” Fierer said. “But we do need to do more research.”
The team plans to investigate the bacterial communities in other cities and to build a continental-scale atlas of airborne bacterial communities, Fierer said. “We don’t know if the patterns we observed in those sites are unique to those cities,” he said. “Does San Francisco have the same bacteria as New York? Nobody knows as yet.”
Fierer believes it is important to pin down the types of bacteria in the air, how these bacteria vary by location and season, and where they are coming from.With this information, scientists can then investigate the possible impacts on human health, he said.
“We need much better information on what sources of bacteria we are breathing in every time we go outside,” Fierer said.
The study was funded by the CIRES Innovative Research Program, the U.S.
Environmental Protection Agency, the National Science Foundation, the Howard Hughes Medical Institute and the National Institutes of Health. The aerosol sample collection for this project was supported by the Lake Michigan Air Directors Consortium.
GUT MICROBES IN HUMANS AND OTHER MAMMALS HEAVILY INFLUENCED BY DIET, SAYS NEW STUDY C U Boulder
May 20th
You are what you eat whether you’re a lion, a giraffe or a human — at least in terms of the bacteria in your gut.
A new study led by the Washington University School of Medicine and involving the University of Colorado Boulder shows gut microbial communities in humans and in a wildly diverse collection of mammals carry out core physiological functions that are heavily influenced by whether they are carnivores, herbivores or omnivores.
The researchers sequenced intestinal microbes in stool samples from 33 mammalian species living in the wild or in zoos in St. Louis and San Diego. In addition to identifying the bacterial species living in the mammalian intestines, they characterized the pool of genes present in each microbial community and their related functions, said senior study author Dr. Jeffrey Gordon of the Washington University School of Medicine in St. Louis.
CU-Boulder professor and study co-author Rob Knight said despite the wide variation of mammals selected for the study, the different gut microbial communities share a set of standard metabolic functions common to all species that play a key role in digestion and immune health. Understanding the variation in human microbial intestine communities holds promise for future clinical research, said Knight, a faculty member in the chemistry and biochemistry department and the computer science department.
A paper on the subject was published in the May 20 issue of Science. Other co-authors on the study included CU-Boulder’s Justin Kuczynski, Dan Knights, Jose Clemente and Antonio Gonzalez, Washington University School of Medicine’s Brian Muegge and Luigi Fontana, and Bernard Henrissat of the Archictecture et Fonction des Macromolecules Biologiques in Marseille, France.
“This was the first time we were able to relate the microbial community members to the specific metabolic functions that were being performed,” said Knight, who also is a Howard Hughes Medical Institute Early Career Scientist. “This is surprising because even members of the same bacterial species can have genomes that are up to 40 percent different in terms of gene content.”
The team extracted DNA from the mammals and humans and used powerful computer methods to sort gene fragments and match them to the DNA of known organisms.
While there was considerable variation of bacterial gut communities between animals, the study showed many of the same microbial genes were found in all of the digestive tracts, with differences in their relative abundance dependent on whether they were meat-eaters, vegetarians or omnivores. Among the mammals whose fecal material was used to sequence gut bacteria microbes included giraffe, bighorn sheep, gazelle, kangaroo, hyena, lion, polar bear, elephant, gorilla, baboon, black bear and squirrel.
The team also showed how diet influences microbial communities in the human gut by sampling 18 lean people who purposely had cut their caloric intake by 25 percent or more using many different dietary strategies. The researchers found that the functions of gut microbes varied according to how much protein the individuals ate, and the bacterial species varied according to how much fiber was consumed.
In a related 2009 study led by CU-Boulder’s Knight, researchers developed the first atlas of microbial diversity across the human body, charting wide variations in microbe populations from the forehead and feet to noses and navels of individuals. One goal of human bacterial studies is to find out what is normal for healthy people, which should provide a baseline for studies looking at human disease states, said Knight, who also is a fellow at CU’s Colorado Initiative for Molecular Biotechnology.
“If we can better understand this microbial variation, we may be able to begin searching for genetic biomarkers for disease,” said Knight. It might someday be possible to identify sites on the human body, including the gut, that would be amenable to microbial community transplants with either natural or engineered microbial systems that would be beneficial to the health of the host, he said.
“Because the human microbiome is much more variable than the human genome, and because it also is much easier to modify, it provides a much more logical starting point for personalized medicine,” he said.
The latest findings emphasize the need to sample humans across the globe with a variety of extreme lifestyles and diets, including hunter-gatherer groups, said the researchers. Such studies could provide insight into the limits of gut bacteria variation and the possibility that human microbes co-evolved with human bodies and cultures, shaping our physiological differences and environmental adaptations.
Members of Knight’s lab and their many collaborators are studying how the human microbiome — all of a person’s hereditary information — is assembled in different people and how it varies in conditions such as obesity, malnutrition and Crohn’s disease. In addition to financial support from HHMI, Knight also has been supported by National Institutes of Health funds to develop new computational tools to better understand the composition and dynamics of microbial communities.
The Science magazine study was supported by the NIH and the Crohn’s and Colitis Foundation of America.
-CU-





















